List of special file formats¶
Contents
Gitools file formats¶
Various formats are accepted in Gitools to represent each of these data types. The following table list the file formats accepted for each data type and in the following sections each file format is described.
| In Gitools | Accepted format | Gitools Expects | Multiple values | Layout |
|---|---|---|---|---|
| Matrix heatmap |
|
|
|
|
| Annotation |
|
|
||
| Module (Gene Sets) |
|
|
Any of the presented formats can be compressed using gzip and recognized if the suffix .gz is appended at the end of the file name, so for example the file matrix.cdm could be compressed using gzip and renamed to matrix.cdm.gz and Gitools would be able to read it without problems.
Import data from external sources¶
Gitools has built-in data importers from different external data sources like Biomart, IntOGen, Ensembl or KEGG. See the importing data section for details.
Gitools also disposes over commands for external control which can be used by other platforms to launch Gitools and launch their own data. See the external control of Gitools if you wish to use the available interfaces.
Prepare your own data¶
The easiest way to create data is using for example a program like Excel or OpenOffice and then export to tabulated text file. See the How to sections with spreadsheet editors.
Continuous data matrix (CDM)¶
CDM file format is a tab delimited matrix of items (i.e. genes) and conditions. The numbers in each cell indicates the values that this item has in that condition. Empty values can be represented with the hyphen ’-’.
It is useful for representing matrices (i.e. expression data for a microarray).
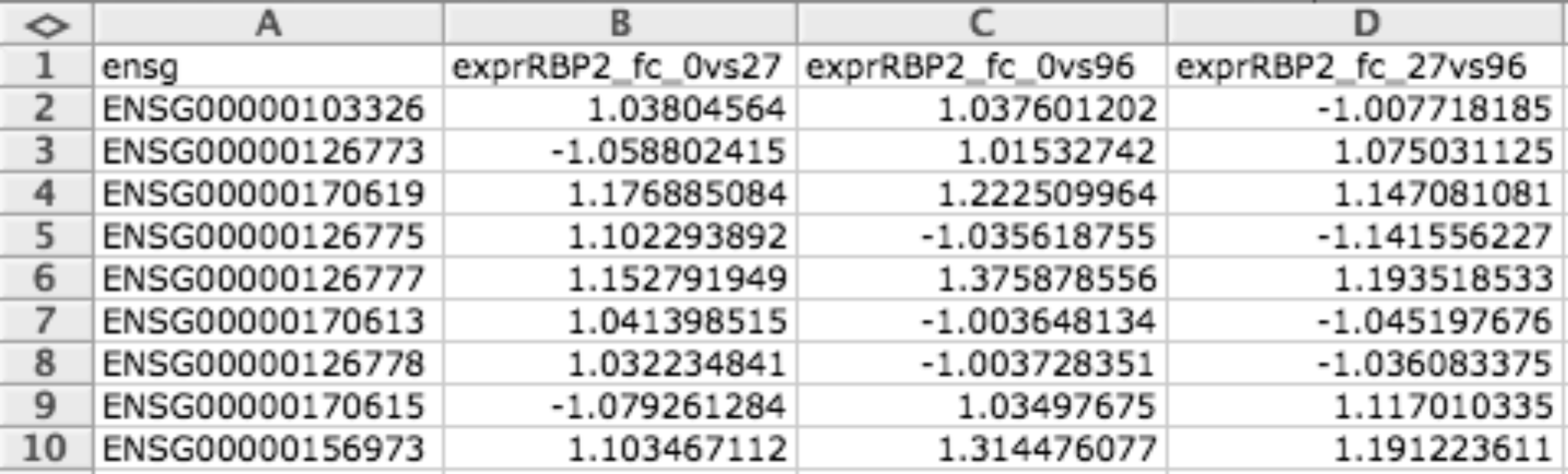
| width: | 700px :align: center
|
|---|
Binary data matrix (BDM)¶
BDM file format is a tab delimited binary matrix of items (i.e. genes) and conditions. Values can only be 1 or 0, and can have different meanings depending what are they used for. Empty values can be represented with the hyphen ’-’.
They are useful for representing matrices as well as modules.
When representing matrices a 1 means that this item (row) presents a positive event (for example a mutation) in this condition (column) and 0 otherwise.
When representing modules, rows correspond to genes or biological elements and each column a different module, then a 1 is used to specify that a given gene or biological element is related to a given module and 0 otherwise.
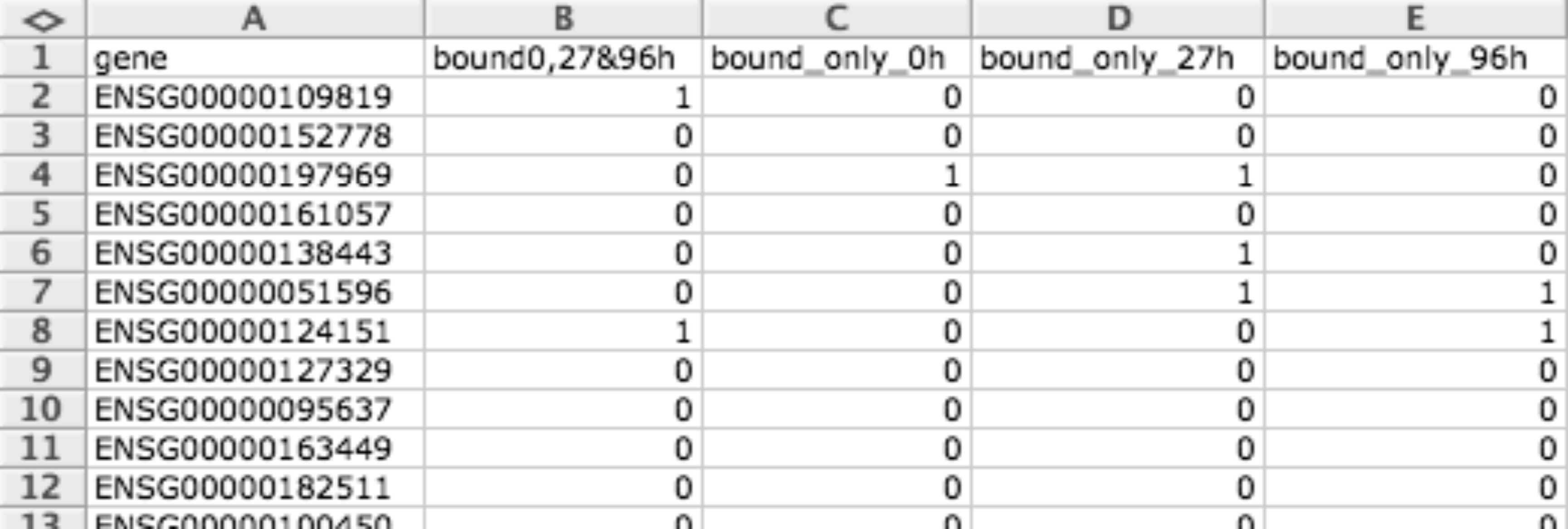
| width: | 700px :align: center
|
|---|
Multi-value data matrix (TDM)¶
TDM file format is a tab delimited file that has contains multiple values per row (gene) and column (sample). The first line is a header line following a line for each cell.
In this following example we see a .tdm-file that contains three columns and two rows.
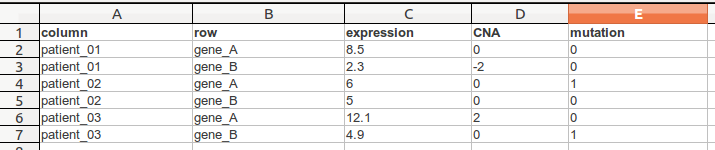
| width: | 700px :align: center
|
|---|
Gene matrix transposed (GMT)¶
GMT file format is a simple tab delimited file to provide gene sets. Each row describes a gene set, the first column indicates the name of the gene set and the second column the description (you can leave description empty), the rest of columns are used to enumerate the genes related to this gene set.
Usually this format is used for representing modules but can also be used to represent binary data matrices (i.e. when you have lists of differentially expressed gens for different conditions).
This format is the same used in GSEA tool .
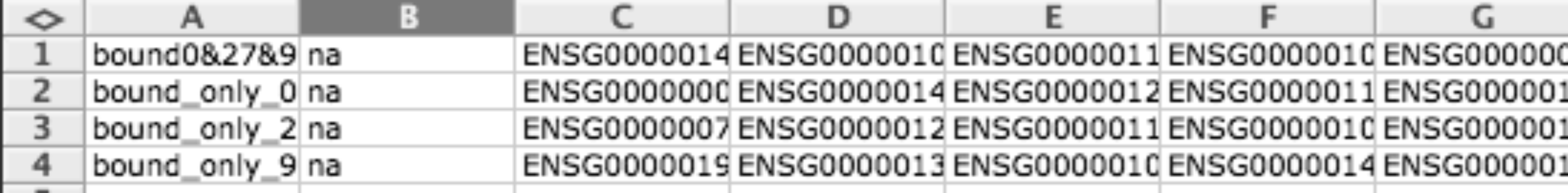
| width: | 700px :align: center
|
|---|
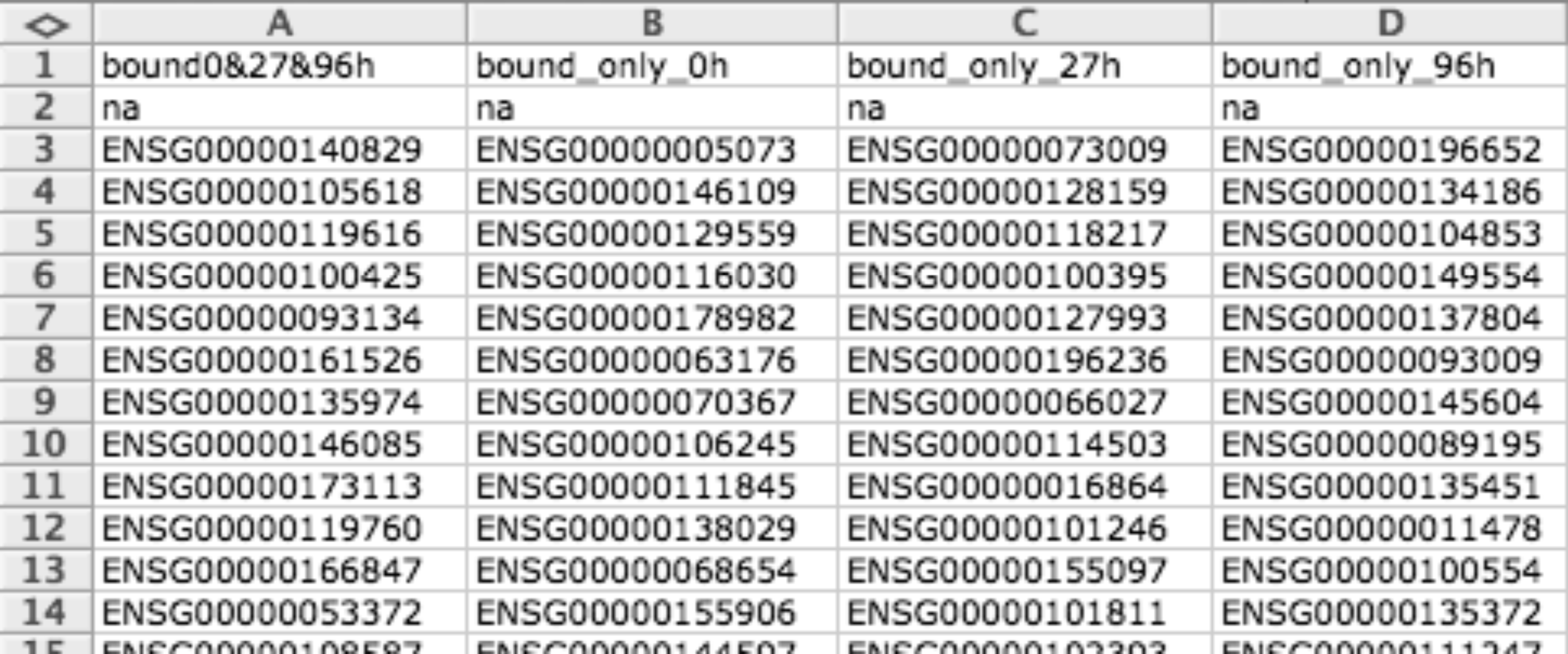
Gene matrix (GMX)¶
GMX file format is a simple tab delimited file to provide gene sets. Each column describes a gene set, the first row indicates the name of the gene set and the second row the description (you can leave description empty), the rest of rows are used to enumerate the genes related to this gene set.
Usually this format is used for representing modules but can also be used to represent binary data matrices (i.e. when you have lists of differentially expressed gens for different conditions).
This format is the same used in GSEA tool .
| width: | 700px :align: center
|
|---|
Two columns mapping (TCM)¶
TCM file format is a simple tab delimited file to provide gene sets. It has two columns, the first column indicates the ID of gene or biological element and the second column the name of the module it belongs to.
Usually this format is used for representing modules but can also be used to represent binary data matrices (i.e. when you have lists of differentially expressed gens for different conditions).
This format uses more disk space than others so it is better to use some of GMX or GMT, but usually it is how you get data from Biomart so Gitools supports it too.
Tab separated values (TSV)¶
This format is a generic text file format used for many different things. The main characteristic is that uses the tab character for separating fields and new line character/s for separating rows. All of the previous formats are based on it.
It can be used to represent matrices, modules and tables.
